JEDLICKA LAB
What we have learned thus far ...
EWS/Fli1-controlled microRNAs as targetable modulators of Ewing Sarcoma pathogenesis
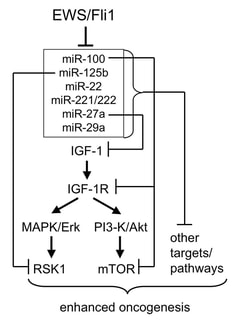
Our investigations began in Ewing Sarcoma. Ewing Sarcoma typically arises in the bone, and less commonly the soft tissues. Ewing Sarcoma is thought to originate in mesenchymal or neural crest progenitor cells, and is the second most common sarcoma at both tissue sites. Ewing Sarcoma pathogenesis is driven by oncogenic proteins arising from chromosomal translocations. The most common such oncoprotein or "oncofusion" is EWS/Fli1. EWS/Fli1 is a hybrid, gain-of-function, transcription factor, which drives Ewing Sarcoma pathogenesis through transcriptional, epigenetic and other mechanisms. In our first series of studies in pediatric sarcoma, we asked how microRNAs regulated by EWS/Fli1 contribute to Ewing Sarcoma pathogenesis, and whether manipulation of specific microRNAs could impact important disease phenotypes.
- McKinsey EL, Parrish JK, Irwin AE, Niemeyer BF, Kern HB, Birks DK and Jedlicka P. A novel oncogenic mechanism in Ewing Sarcoma involving IGF pathway targeting by EWS/Fli1-regulated microRNAs. Oncogene (2011). https://pubmed.ncbi.nlm.nih.gov/21643012
- Dylla L and Jedlicka P. Growth-promoting role of the miR-106a~363 cluster in Ewing Sarcoma. PLoS One (2013). https://pubmed.ncbi.nlm.nih.gov/23638178
- Robin TP, Smith A, McKinsey E, Reaves L, Jedlicka P and Ford HL. EWS/Fli1 regulates Eya3 in Ewing's sarcoma via modulation of microRNA-708, resulting in increased cell survival and chemoresistance. Molecular Cancer Research (2012). https://pubmed.ncbi.nlm.nih.gov/22723308
- Niemeyer BF, Parrish JK, Spoelstra NS, Joyal T, Richer JK and Jedlicka P. Variable expression of PIK3R3 and PTEN in Ewing Sarcoma impacts oncogenic phenotypes. PLoS One (2015). https://pubmed.ncbi.nlm.nih.gov/25603314
- Moore C, Parrish JK and Jedlicka P. MiR-193b, downregulated in Ewing Sarcoma, targets the ErbB4 oncogene to inhibit anchorage-independent growth. PLoS ONE (2017). https://pubmed.ncbi.nlm.nih.gov/28542597
Reviews:
- Jedlicka P. Ewing Sarcoma, an Enigmatic Malignancy of Likely Progenitor Cell Origin, Driven by Transcription Factor Oncogenic Fusions. International Journal of Clinical and Experimental Pathology (2010). https://pubmed.ncbi.nlm.nih.gov/20490326
- Dylla L, Moore C and Jedlicka P. MicroRNAs in Ewing Sarcoma. Frontiers in Oncology (2013). https://pubmed.ncbi.nlm.nih.gov/23543617
Biology and targeting of Jumonji-domain histone demethylases in Ewing Sarcoma progression
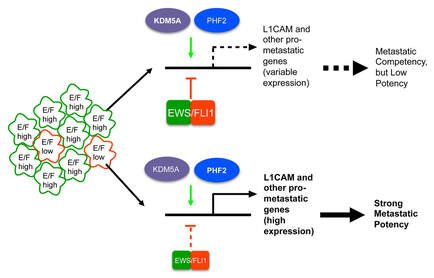
Our studies of microRNAs also led us to interesting, previously unexplored and targetable, epigenetic regulators in Ewing Sarcoma, namely KDM3A and other histone demethylases belonging to the Jumonji-domain family (JHDMs). We found that members of the JHDM family exert potent disease-promoting properties in Ewing Sarcoma. During our initial investigations of JHDMs, data from other laboratories emerged showing that EWS/Fli1 exerts mixed, including inhibitory, effects on Ewing Sarcoma metastatic properties. Interestingly, our findings showed that some of the disease-promoting effects of JHDMs in Ewing Sarcoma act in opposition to the metastasis-repressive effects of EWS/Fli1. We also demonstrated activity of a pan-JHDM pharmacologic inhibitor (JIB-04) in Ewing Sarcoma.
- Parrish JK, Sechler M, Winn RA and Jedlicka P. The histone demethylase KDM3A is a microRNA-22-regulated tumor promoter in Ewing Sarcoma. Oncogene (2015). https://pubmed.ncbi.nlm.nih.gov/24362521
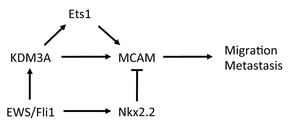
- Sechler M, Parrish JK, Birks DK and Jedlicka P. The histone demethylase KDM3A, and its downstream target MCAM, promote Ewing Sarcoma cell migration and metastasis. Oncogene (2017). https://pubmed.ncbi.nlm.nih.gov/28319067
- Parrish JK, McCann TS, Sechler M, Sobral LM, Ren W, Jones KL, Tan AC and Jedlicka P. The Jumonji-domain histone demethylase inhibitor JIB-04 deregulates oncogenic programs and increases DNA damage in Ewing Sarcoma, resulting in impaired cell proliferation and survival, and reduced tumor growth. Oncotarget (2018). https://pubmed.ncbi.nlm.nih.gov/30237855
- McCann TS, Parrish JK, Hsieh J, Sobral LM, Self C, Sechler M, Jones KL, Goodspeed A, Costello J and Jedlicka P. KDM5A and PHF2 positively control expression of pro-metastatic genes repressed by EWS/Fli1, and promote growth and metastatic properties in Ewing Sarcoma. Oncotarget (2020). https://pubmed.ncbi.nlm.nih.gov/33196691
Reviews:
- Jedlicka P. The potential of KDM3A as a therapeutic target in Ewing Sarcoma and other cancers. Expert Opinion on Therapeutic Targets (2017). https://pubmed.ncbi.nlm.nih.gov/29022407
- McCann TS, Sobral LM, Self C, Hsieh J, Sechler M and Jedlicka P. Biology and targeting of the Jumonji-domain histone demethylase family in childhood neoplasia: a preclinical overview. Expert Opinion on Therapeutic Targets (2019) https://pubmed.ncbi.nlm.nih.gov/30759030
KDM3A/Ets1 epigenetic axis in Rhabdomyosarcoma progression
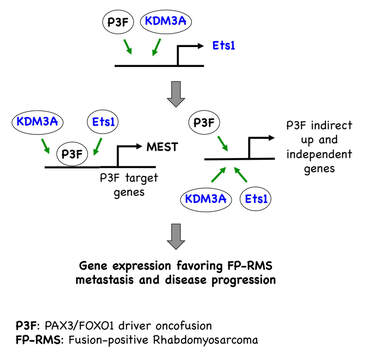
Early on in our studies of KDM3A in Ewing Sarcoma, we noted that this epigenetic regulator was also highly expressed in Rhabdomyosarcoma (RMS). RMS is a sarcoma with skeletal muscle differentiation, is the most common cancer of soft tissue in children, and mainly exists as two different disease sybtypes. Fusion-negative RMS (FN-RMS) is molecularly heterogeneous, with frequent mutations in receptor tyrosine kinase (RTK)/Ras/PI3K signaling axes. Fusion-positive RMS (FP-RMS), as its name implies, is driven by oncogenic proteins arising from chromosomal translocations, similar to Ewing Sarcoma. The most common driver oncofusion in FP-RMS is PAX3/FOXO1, and less commonly PAX7/FOXO1. Similar to Ewing Sarcoma, PAX3/7-FOXO1 oncofusions are hybrid, gain-of-function, transcription factors that drive FP-RMS pathogenesis through transcriptional and epigenetic mechanisms. Our studies thus far point to important roles of KDM3A, and the downstream transcription factor Ets1, in the aggressive properties and disease progression of both RMS subtypes, and especially FP-RMS.
- Sobral LM, Sechler M, Parrish JK, McCann TS, Jones KL, Black JC and Jedlicka P. KDM3A/Ets1/MCAM axis promotes growth and metastatic properties in Rhabdomyosarcoma. Genes and Cancer (2020). https://pubmed.ncbi.nlm.nih.gov/32577157
- Sobral LM, Hicks HM, Parrish JK, Hsieh J, McCann TS, Goodspeed A, Costello J, Black JC and Jedlicka P. KDM3A/Ets1 epigenetic axis contributes to PAX3/FOXO1-driven and independent disease-promoting gene expression in fusion-positive Rhabdomyosarcoma. Molecular Oncology (2020).https://pubmed.ncbi.nlm.nih.gov/32697014
Our investigations in pediatric sarcoma have been supported by: